Genomic Solutions for Health
.webp/:/cr=t:0%25,l:0%25,w:100%25,h:100%25/rs=w:365,h:365,cg:true)
Advanced Genomic Analysis
AI Driven Drug Discovery & Development
Transcriptomics & Single-Cell Analysis
Whole Genome Sequencing (WGS) and Exome Sequencing (WES) provide comprehensive genomic analysis, including variant calling for SNPs, Indels, CNVs, and SVs, along with clinical annotation through advanced bioinformatics techniques. Targeted Sequencing allows for the analysis of custom assay design using both custom and pre-defined gene pan
Whole Genome Sequencing (WGS) and Exome Sequencing (WES) provide comprehensive genomic analysis, including variant calling for SNPs, Indels, CNVs, and SVs, along with clinical annotation through advanced bioinformatics techniques. Targeted Sequencing allows for the analysis of custom assay design using both custom and pre-defined gene panels tailored for specific disease areas. In the realm of Cancer Genomics, our services include tumor-normal pair analysis, somatic variant detection, and assessment of tumor mutational burden (TMB), all of which are crucial for drug discovery.

Transcriptomics & Single-Cell Analysis
AI Driven Drug Discovery & Development
Transcriptomics & Single-Cell Analysis
Single-Cell RNA-Seq (scRNA-seq) enables advanced bioinformatics applications such as cell-type clustering, trajectory analysis, differential expression, and the identification of rare cell populations, which are crucial in cancer genomics. In addition, Bulk RNA-Seq Analysis focuses on differential gene expression (DGE), pathway enrichment
Single-Cell RNA-Seq (scRNA-seq) enables advanced bioinformatics applications such as cell-type clustering, trajectory analysis, differential expression, and the identification of rare cell populations, which are crucial in cancer genomics. In addition, Bulk RNA-Seq Analysis focuses on differential gene expression (DGE), pathway enrichment, and gene-set enrichment analysis (GSEA), providing insights for drug discovery and genomic analysis. Furthermore, Spatial Transcriptomics allows for the mapping of gene expression patterns within the morphological context of tissue, enhancing our understanding of targeted sequencing and whole genome sequencing.

AI Driven Drug Discovery & Development
AI Driven Drug Discovery & Development
AI Driven Drug Discovery & Development
Target Identification & Validation: Leveraging bioinformatics and multi-omics data to pinpoint promising new drug targets in cancer genomics. Biomarker Discovery: Utilizing genomic analysis to identify predictive and prognostic biomarkers for effective patient stratification. Mechanism of Action (MoA) Studies: Elucidating how therapeutic
Target Identification & Validation: Leveraging bioinformatics and multi-omics data to pinpoint promising new drug targets in cancer genomics. Biomarker Discovery: Utilizing genomic analysis to identify predictive and prognostic biomarkers for effective patient stratification. Mechanism of Action (MoA) Studies: Elucidating how therapeutic compounds work at a molecular level through custom assay design, including targeted sequencing and whole genome sequencing.

Cytogenetics Analysis & Reporting
Custom Assay Design & Bioinformatics Pipelines
AI Driven Drug Discovery & Development
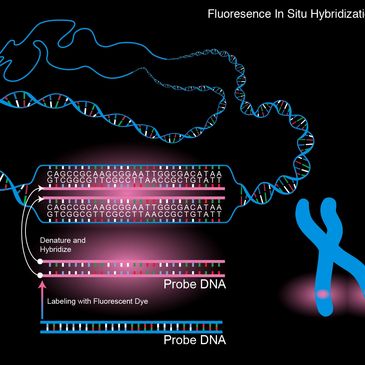
Karyotyping Analysis: Utilizing bioinformatics for digital analysis and annotation of chromosomal banding in both constitutional and cancer genomics. FISH Analysis: This includes quantitative analysis and reporting for Fluorescence In Situ Hybridization, covering fusion, break-apart, and copy number probes to aid in targeted sequencing. C
Karyotyping Analysis: Utilizing bioinformatics for digital analysis and annotation of chromosomal banding in both constitutional and cancer genomics. FISH Analysis: This includes quantitative analysis and reporting for Fluorescence In Situ Hybridization, covering fusion, break-apart, and copy number probes to aid in targeted sequencing. Chromosomal Microarray (CMA) Analysis: An in-depth genomic analysis of microarray data to detect microdeletions, duplications, and copy-neutral LOH, which is essential in custom assay design and drug discovery.

Expert Consulting & Troubleshooting Support
Custom Assay Design & Bioinformatics Pipelines
Custom Assay Design & Bioinformatics Pipelines
NGS Experimental Design: Get expert advice on choosing the right sequencing platform, including targeted sequencing and whole genome sequencing, as well as custom assay design and strategy for your clinical research question in cancer genomics. Molecular Assay Analysis: Perform genomic analysis with quantitative assessment of FISH (Fluore
NGS Experimental Design: Get expert advice on choosing the right sequencing platform, including targeted sequencing and whole genome sequencing, as well as custom assay design and strategy for your clinical research question in cancer genomics. Molecular Assay Analysis: Perform genomic analysis with quantitative assessment of FISH (Fluorescence In Situ Hybridization) and qPCR (Quantitative PCR) data. Wet-Lab Protocol Troubleshooting: Receive one-on-one support to optimize or troubleshoot key molecular assays, including NGS library preparation, qPCR, and FISH, all essential for drug discovery.
Custom Assay Design & Bioinformatics Pipelines
Custom Assay Design & Bioinformatics Pipelines
Custom Assay Design & Bioinformatics Pipelines
Custom Bioinformatics Pipeline Development: We create automated, scalable, and clinically-validated bioinformatics pipelines tailored to your specific research or diagnostic workflow in cancer genomics. qPCR Assay Design: Viral Load: Highly sensitive probes for quantitative viral load monitoring. Genotyping: Allele-specific primers and pr
Custom Bioinformatics Pipeline Development: We create automated, scalable, and clinically-validated bioinformatics pipelines tailored to your specific research or diagnostic workflow in cancer genomics. qPCR Assay Design: Viral Load: Highly sensitive probes for quantitative viral load monitoring. Genotyping: Allele-specific primers and probes for SNP and Indel detection. Gene Expression: Custom assay design includes TaqMan and SYBR Green assays for quantifying target gene expression. FISH Probe Design: In silico design and validation of custom probes for Fluorescence In Situ Hybridization, targeting specific gene loci, translocations, or chromosomal regions. Custom NGS Panel Design: We design, validate, and optimize targeted sequencing panels (amplicon or hybrid-capture) focused on your specific genes of interest for any clinical application, supporting drug discovery and genomic analysis through whole genome sequencing.
Copyright © 2025 Clinx Genomics - All Rights Reserved.
This website uses cookies.
We use cookies to analyze website traffic and optimize your website experience. By accepting our use of cookies, your data will be aggregated with all other user data.